UV Resistance and Photoreactivation of Extremophiles from High-Altitude Andean Lakes
Virginia H. Albarracín1,2,3, Wolfgang Gärtner3, María E. Farías1
1Lab of Microbial Research on Andean Lakes (LIMLA)
Pilot Plant of Microbial Industrial Processes (PROIMI)
National Research Council of Argentina (CONICET)
Av. Belgrano y Pasaje Caseros, (4000) Tucumán, Argentina.
virginia@proimi.org.ar
mefarias@proimi.org.ar
2Natural Sciences College and Miguel Lillo Institute
National University of Tucumán. Miguel Lillo 205
(4000) Tucumán, Argentina
3Max-Planck-Institute for Chemical Energy Conversion
Stiftstrasse 34-36, D-45470 Mülheim, Germany
virginia-helena.albarracin@cec.mpg.de
wolfgang.gaertner@cec.mpg.de
High-Altitude Andean Lakes (HAAL): A Diverse Source of Poly-Extremophilic Microorganisms
Anthropomorphically, an extreme environment is one in which physical conditions are not conducive for human life. On the other hand, for many species extreme environments are the norm. Thus, we define these habitats as those that experience steady or fluctuating exposure to one or more environmental factors, such as salinity, osmolarity, desiccation, ultraviolet (UV) radiation, barometric pressure, pH, or temperature (Seufferheld et al., 2008).
Microorganisms that colonize extreme environments are called extremophiles. This group includes representatives of all three domains (Bacteria, Archaea, and Eukarya); they are categorized into subgroups according to the specific environmental characteristics of their habitats, e.g., psychrophilic, thermophilic, halophilic, alkalophilic, or acidophilic.
Extreme environments have been subject to intensive studies, focusing attention on the diversity of organisms and molecular and regulatory mechanisms involved. The products obtainable from extremophiles, such as proteins, enzymes ("extremozymes") and compatible solutes, are of great interest to biotechnology (Sanchez et al., 2009). Past examples include biochemicals used for detergent formulations, leather and paper processing, biofuels, bioremediation, UV-blocking, and new antibiotics. However, potentially beneficial biomolecules still remain to be discovered from unexplored extreme environments. Not to forget; this field of research has also attracted attention because of its impact on the studies of possible existence of life on other planets.
The High-Altitude Andean Lakes (HAAL) at the dry central Andes region in South America represent an almost unexplored ecosystem of shallow lakes and salterns at altitudes of 3,000–6,000 m above sea level (asl), extended across political boundaries of the countries of Argentina, Chile, Bolivia and Peru. Our research group has mainly focused on the lakes and salterns present in the Argentinean side (Figure 1), and we could determine their abundant and diverse microbiota. Though difficult to explore, these ecosystems present a number of remarkable properties for the study of extreme biological systems: (i) they are pristine and isolated with no access roads; (ii) they are distant from each other (more than 500–700 km); (iii) they are surrounded by desert; (iv) they are the habitat of enormous populations of three flamingos species that migrate among these wetlands, and act as microbial dispersers; (v) they are located in a geographical area where the solar irradiation, and thus the UV exposure, is the highest on Earth; (vi) they are subject to daily large temperature fluctuations (up to 20º C of difference between day and night); (vii) they display high salinity and extremely high arsenic content (of geochemical origin); (viii) they are located in a volcanic landscape; hydrothermal vents are common inputs to the lakes modifying the nutrients flow and water temperature. These outstandingly hostile environmental conditions are considered to significantly resemble those of the Earth's early atmosphere, as has been stated by NASA (Cabrol et al., 2007; Farias et al., 2009).
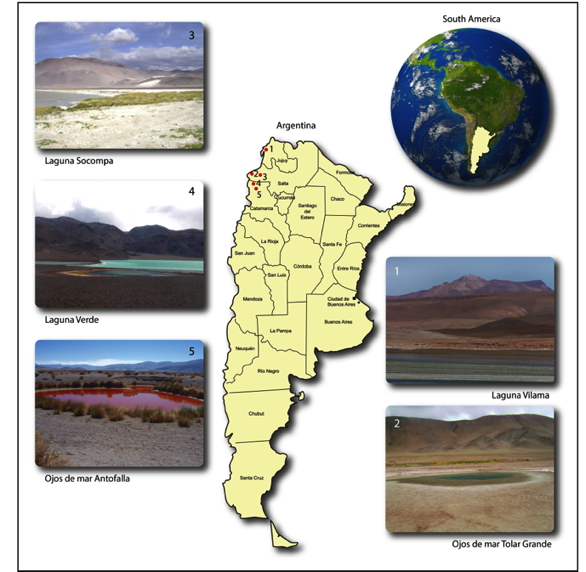
Figure 1. Geographic location of some lakes sampled at the HAAL, in which an abundant microbiodiversity of extremophiles was encountered: Laguna Vilama (4,500 m) in Jujuy, Laguna Socompa (3,750 m) and Ojos de Mar Tolar Grande (3,510 m) in Salta, Laguna Verde (4,100 m) and Ojos de Mar Antofalla (3,350 m) in Catamarca, all of them located in Argentina.
Although exposed to extreme conditions, an outstanding microbial biodiversity has developed in most ecotopes: water, sediments, soil, fumaroles, bioevaporites, microbial mats, and even in multi-layered flat mats and stubby pillars called stromatolites (among all sites where stromatolites are described, those found in the HAAL are located at the highest altitudes on Earth) (Farias et al., 2011, 2013). In fact, after a decade of research expeditions at the HAAL, a collection of extremophiles, covering ~500 strains, is available (Ordoñez et al., 2009). This worldwide-open collection harbors prokaryotes as well as eukaryotes able to resist several extreme conditions (alkalinity, hypersalinity, UV-A(-B) radiation). HAAL microbial diversity is not only of worldwide interest as an exceptional template for early life development; their indigenous extremophiles are a treasure chest of biotechnological importance; "genome mining" can unravel exceptional biomolecules (i.e., extremoenzymes) for industrial or biomedical applications, even as their characterization is still at its beginning (Albarracin and Farias, 2012).
Ultraviolet Radiation Resistance as The Rule For HAAL'S Microbes.
Ultraviolet radiation is part of the solar electromagnetic spectrum, and it is an essential factor for many global biological and environmental phenomena. There are three major subtypes of UV rays, namely, UV-A (320–400 nm), UV-B (280–320 nm) and UV-C (100–280 nm). UV-B and UV-C are detrimental to life because of the strong absorption of wavelengths below 320 nm by DNA molecules, whereas UV-A (320–400 nm) causes mainly indirect damage to DNA, proteins, and lipids through reactive oxygen intermediates that are generated by photosensitizing molecules showing an absorption band in this wavelength range. UV-A accounts for about 95% of the total UV energy that reaches the Earth‘s surface, the remaining 5% being UV-B, as shorter wavelengths are increasingly absorbed by the atmosphere; UV-C gets totally absorbed by stratospheric gases, mainly oxygen and ozone, and thus fails to reach the troposphere. Furthermore, since ozone molecules very effectively screen out UV-B, only a small fraction actually reaches the surface, contrary to most of UV-A (for more information, see the Modules on UV Radiation Photobiology).
The biological consequences arising from increased UV irradiance are numerous. In terrestrial ecosystems, these affect plants, pathogens, herbivores, soil microbes. As each type of organism reacts to induced UV damage in a different manner, the eventual changes in balance can possibly lead to significant alterations in carbon and nitrogen cycling. Furthermore, apart from ozone concentration dependence, UV irradiance is also affected by climate change factors, thus complex interactions are expected to occur, thereby diversely affecting terrestrial ecosystems.
Exposure to UV radiation is considered to be especially harmful to microorganisms, because they have haploid genomes with little or no functional redundancy. In addition, they are small and lack thick, protective cell walls (Ponder et al., 2005). Nevertheless, some microorganisms have evolved complex mechanisms that make them resistant to large doses of radiation; the most extensively studied so far is Deinococcus radiodurans, a vegetative, easily cultured, and nonpathogenic bacterium belonging to the family Deinococcaceae (Battista et al., 1999; Makarova et al., 2001). This aerobic, large (1 to 2-mm) tetrad-forming soil bacterium is best known for its supreme resistance to ionizing radiation, but also it is resistant towards UV radiation, hydrogen peroxide, and numerous other agents that damage DNA (Minton, 1994; Lange et al., 1998, Battista et al., 1999), as well as being highly resistant to desiccation (Mattimore and Battista, 1996). It can survive acute exposures to gamma radiation that exceed 1,500 krads without dying or undergoing induced mutation (Daly et al., 1994), while an acute dose of only 100 to 200 kilorads sterilize a culture of Escherichia coli (Lange et al., 1998). In exponential phase, D. radiodurans is 33-fold more resistant to UV than is E. coli (Sweet and Moseley, 1974). Furthermore, survivors of extreme ionizing radiation, UV, or bulky chemical-adduct exposures do not show any mutagenesis greater than that occurring after a single round of normal replication (Sweet and Moseley, 1974; 1976). Supporting this outstanding resistance phenotype, several mechanisms have been described: i.e., chromosomal multiplicity, repair pathways that include excision repair, mismatch repair, and recombinational repair (for a detailed review, see Makarova et al., 2001), as well as a very efficient recA-independent single-stranded DNA annealing repair pathway, which is active during and immediately after DNA damage, and before the onset of recA-dependent repair (Daly and Minton, 1996).
The effects of UV radiation on aquatic systems have been thoroughly studied in marine environments (Joux et al., 1999; Agogue et al., 2005; Hernandez et al., 2007), indicating that this radiation eventually affects the whole community, having an impact on photosynthesis, biomass production, and the community composition (Winter et al., 2001; Banaszak, 2003). Less attention has been given to studies on the impact of UV radiation on bacterioplankton from other aquatic systems, such as mountain lakes (Winter et al., 2001). Several reports have been presented on the effects of solar UV irradiation on plankton from alpine lakes (Williamson et al., 1995; Halac et al., 1997; Winter et al., 2001; Häder et al., 2007). A few studies on biodiversity in the Himalayas have been presented (Liu et al., 2006; Jiang et al., 2007).
Due to the high altitude and the geographical and physicochemical characteristics of HAAL, UV radiation is one of the most limiting abiotic factors for HAAL microbial communities. Solar irradiance is much higher than at sea level, with instantaneous UV-B flux reaching 17 W m-2 in some lakes (compared with 0.1-0.4 W m-2 at sea level). In accordance with this, HAAL-isolated strains display an intrinsic and high UV-B resistance. In many cases, UV-B irradiation was used as a pre-treatment for sample preparation before the isolation procedure takes place, which allows to differentially isolate the most UV-B-resistant strain from a particular sample (Fernandez-Zenoff et al., 2006; Zenoff et al., 2006). Following this procedure a set of almost one hundred UV-B-resistant strains were isolated (Figure 2), characterized and identified as belonging to diverse taxonomic groups (Table 1; Fernandez-Zenoff et al., 2006; Zenoff et al., 2006; Dib et al., 2008; Ordoñez et al., 2009; Flores et al., 2009; Bequer-Urbano et al., 2013).
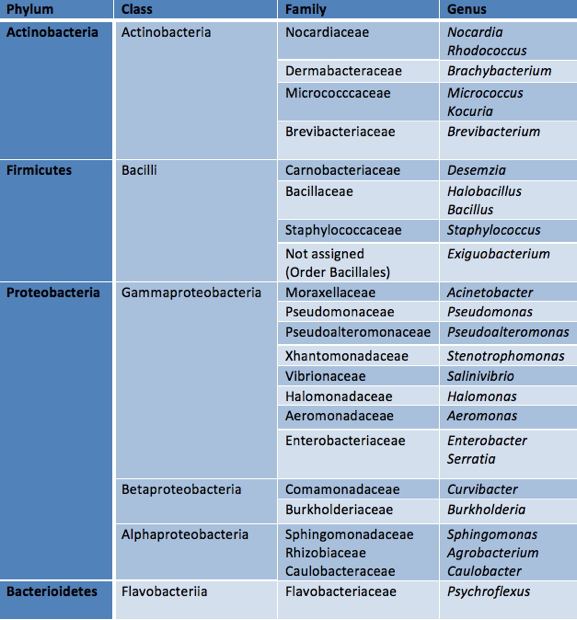
Table 1. Taxonomic affiliations of UV-B-resistant bacteria isolated from High-Altitude Andean Lakes (HAAL).

Figure 2. Some of HAAL UV-B-resistant bacteria observed under a stereoscopic lamp (magnification for the left image is 3X, and for the other two is 0.8X). From left to right: Rhodococcus sp. A5, Stenotrophomonas sp. N50, Pseudomonas sp. V1.
Most UV-B-resistant bacteria from HAAL belong to the genus Acinetobacter, gram-negative, oxidase-negative, non-motile, and strictly aerobic bacteria, which tend to grow as paired cocci (Towner, 2009). Acinetobacter clinical isolates have received growing attention due to the clinical relevance of their multi-drug resistance, but less consideration has been given to environmental isolates from Acinetobacter, despite their metabolic versatility, biotechnological potential, and their well-known wide resistance to environmental stress (Abdel-El-Haleem, 2003). The Acinetobacter sp. strains from the HAAL turned out to be even more resistant than their closest taxonomical neighbors: A. baumannii DSM 30007, A. johnsonii DSM 6963 and Acinetobacter lwoffii DSM 2403 (Fernandez Zenoff et al., 2006; Albarracin et al., 2012; Di Capua et al., 2012). To highlight the UV-B resistance strength of HAAL strains, we compared the survival and photoreactivation (see module on Photoreactivation) after UV-B exposure of Acinetobacter sp. Ver3 with an E. coli strain (Figure 3), since extensive research has been done on this species.
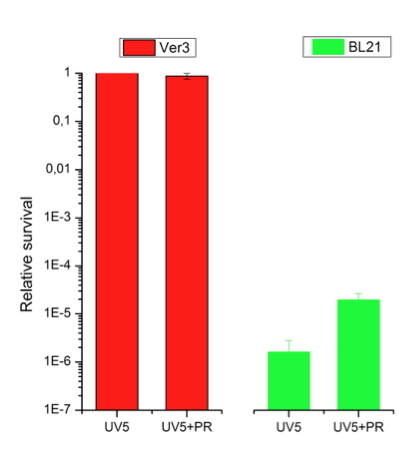
Figure 3. Survival and photoreactivation of Acinetobacter sp Ver3 (HAAL strain) after 5 min of UV-B irradiation (1.2 kJ m-2, Vilbert Lourmat VL-4; λmax at 312 nm with an average intensity of 0.387 mW cm-2) in comparison with an E. coli strain [BL21 (Stratagene)]. UV5: cell suspensions were exposed to 5 min of UV-B irradiation and immediately transfered to agar plates, and incubated in the dark for 24 h. UV5+PR: cell suspensions were exposed to 5 min of UV-B irradiation, plus 120 min of photoreactivating light, and then transfered to agar plates, and incubated in the dark for 24 h. The results are shown as relative values with respect to non-photoreactivated cells.
-B The treatment called UV5 let us observe the recovery of cells based on protection and/or repair of UV-B-induced photodamage in the dark, i.e., without photoreactivation, while the treatment, UV5+PR, means that the cells where photoreactivated prior to incubation in the dark. At the same UV-B dose tested (1.2 kJ m-2), Ver3 cells showed no changes in survival, while the E. coli strain showed a drastic depletion of the bacterial population of five or more orders of magnitude (Figure 3). This was observed even if cells were photoreactivated prior to incubation in the dark. For the E. coli strain, UV-B exposure was more harmful when cells were not photoreactivated before incubation (Figure 3).
On the other hand, the survival of Ver3 cells was 100% under both treatments, indicating that it displays efficient mechanisms to protect from or counteract act the photodamage in the dark, which may be of similar efficiency to the mechanisms acting when light is available for 120 min prior to incubation in the dark (Figure 3). Three mechanisms are expected to be the basis behind the HAAL strains high UV-B-resistance profile: i) competent protection against reactive oxygen species (ROS), potentially damaging agents for lipids, proteins and nucleic acids; ii) capability of bypassing DNA lesions, and iii) efficient repairing of DNA photoproducts. A great number of specific and highly conserved DNA repair mechanisms has been developed against DNA damage; these are photoreactivation (see below), excision repair, recombination repair, mismatch repair (MMR), and double-strand break (DSB) repair. In addition, damage tolerance (dimer bypass), SOS (save our soul) response, checkpoint activation, and programmed cell death (PCD) or apoptosis efficiently act against DNA lesions, ensuring the genomic integrity (for more information, see Modules on UV Radiation Photobiology).
The first topic listed (protection), is accomplished by Acinetobacter sp. Ver3 and Ver7 (both isolates from Lake Verde, at 4.100 m asl) that both displayed unusually high catalase activity with or without UV-B-challenge (Di Capua et al., 2012). Moreover, inhibition of catalase by 3-amino-1,2,4-triazole (AT) resulted in a decrease of the observed tolerance to UV-B radiation by Ver7, suggesting the involvement of this enzyme in the resistance against UV-B radiation.
Although HAAL strains displayed higher UV-B resistance profiles than control strains, they also displayed the greatest accumulation of photoproducts after 10 min of UV-B irradiation, roughly 25% more photoproducts than the controls (A. baumannii DSM 30007, A. johnsonii DSM 6963) (Albarracin et al., 2012). This may suggest that the ability of UV-B resistance is not strictly related to a lower DNA photodamage. This pattern of high survival together with a high photoproduct accumulation could alternatively be explained by a DNA lesion bypass, a typical mechanism of error-prone dark repair (Sinha and Häder, 2002), which keeps the cells active with DNA replication (by polymerase V) without DNA damage repair (Fernandez-Zenoff et al., 2006; Albarracin et al., 2012).
As a third mechanism to counteract the detrimental effects of UV-B radiation, HAAL strains diminish the CPD (cyclobutane pyrimidine dimer) content more efficiently than control strains not isolated from those environments (Fernandez-Zenoff et al., 2006; Albarracin et al., 2012). This outstanding UV-B resistance, as well as their efficient photorepairing ability is a strong hint for the presence of highly active photolyases in these novel extremophilic strains.
Photolyases and Cryptochromes of Extreme Bacteria From the HAAL
As HAAL are ecosystems highly irradiated by light, we expected to find a wide occurrence and rich diversity of photoreceptors in their extremophiles, being the ones able to act in DNA damage repairing as the most important factors for understanding their survival mechanisms.
Photolyases (PL) are monomeric flavoproteins of 53-66 kDa, which, depending on the type of organism, contain between 450 and 620 amino acids, and a non-covalently bound flavin adenine dinucleotide (FAD) as cofactor in a 1:1 ratio. In addition, they also carry an antenna pigment, such as deazaflavin or methenyltetrahydrofolate derivatives (Sancar, 2003).
Photolyases are efficient enzymes repairing DNA photoproducts, namely the CPD or the (6-4) pyrimidine–pyrimidone photoproduct [(6-4)photoproduct] in single-stranded DNA (ssDNA) as well as in double-stranded DNA (dsDNA). For this function, photolyases bind tightly to the photoproduct in the dark, and are activated by the absorption of UV-A or blue light by the reduced form of FAD (FADH-). This electron transfer process, in turn, starts a downhill electron-transfer-driven reaction affording the excision of the two C–C bonds in the CPD, and the reformation of two monomeric pyrimidine bases (Weber, 2005).
The most extensively studied photorepair mechanism among bacteria is that of the E. coli photolyase, belonging to the Class I CPD-photolyases for which also the crystal structure (Park et al., 1995), DNA binding site and the CPD to DNA monomer equilibrium have been determined (Brettel and Byrdin, 2000; Sancar, 2000). In this group, there are also other crystallized CPD-photolyases from the cyanobacterium Anacystis nidulans (Tamada et al., 1997) , the eubacterium Thermus thermophilus (Klar et al., 2006), and the archaeon Sulfolobus tokodaii (Fujihashi et al., 2007). Detailed work has also been performed on the energetics of radical transfer in DNA photolyase and the photoreaction of the photolyases from Anacystis nidulans, where a radical reaction chain involving tyrosine and tryptophan residues has been identified (Tamada et al., 1997).
Photolyases together with the cryptochromes (Cry) form a divergent enzyme family [hereafter, CPF(Cryptochrome-Photolyase Family)] occurring in all three kingdoms of life, most likely because of the need of early forms of life on Earth to counter act DNA damage caused by UV radiation (Oberpichler et al., 2011). They share strong sequence and structural similarity, making a common ancestor probable for both these protein classes. Typically, cryptochromes are involved in circadian clock entrainment, stomatal cell opening, and many other regulating processes (Cashmore, 2003). The discrimination between photolyases and cryptochromes became recently blurred, as cryptochromes, such as the DASH-type (Drosophila-Arabidopsis-Synecoccochus, Homo sapiens) or CryA from Aspergillus nidulans, exert dual functions by being competent in signaling and DNA repair. Both, photolyases and cryptochromes display a bilobal architecture featuring an amino (N)-terminal Rossmann fold, and a carboxy (C)-terminal catalytic domain. The catalytic cofactor, flavin adenine dinucleotide (FAD), is bound in a U-shaped conformation, such that the adenine part is in close contact with the isoalloxazine ring moiety. In photolyases, the FAD chromophore adopts all three possible oxidation states, and is indispensable for catalytic or signaling processes. The N-terminus might bind antenna chromophores to enhance absorbance cross-sections towards blue light.
According to their functions, CPF proteins can be divided into three major classes: CPD photolyases, (6-4) photolyases and Cry. Phylogenetically, the groups are further divided into the class I to class III CPD photolyases, plant Cry, DASH Cry, as well as animal type I- and type II-Cry, respectively. Animal Cry are closely related to (6-4) photolyases, while plant Cry form a sister group of the class III CPD photolyases (Oberpichler et al., 2011). Proteobacteria and cyanobacteria harbor a new class of cryptochromes, called CryPro. A crystal structure is already available for one of its members, cryptochrome B (CryB) from Rhodobacter sphaeroides, where this protein is involved in the regulation of photosynthesis gene expression (Geisselbrecht et al., 2012). The structure reveals that, in addition to the photolyase-like fold, CryB contains two cofactors only conserved in the CryPro subfamily: 6,7-dimethyl-8-ribityl-lumazine in the antenna-binding domain and a [4Fe-4S] cluster within the catalytic domain. The latter closely resembles the iron–sulphur cluster harboring part of the large subunit of PriL primase, indicating that PriL is evolutionarily related to the CryPro class of cryptochromes.
Mining in recently available genomes of three UV-B-resistant bacteria Acinetobacter sp. Ver3, Exiguobacterium sp. S17, and Nesterenkonia sp. Act20 isolated from the shallow water of Laguna Verde (4,400 m asl), modern stromatolites, and surrounding soil of Laguna Socompa (3,750 m asl) (Figure 1), respectively, confirmed the presence of a rich diversity of photoreceptors from the CPF family (Table 2) in the HAAL isolates (Figure 4).
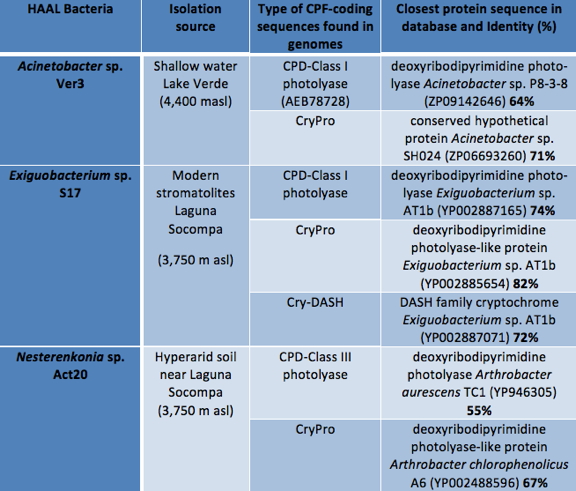
Table 2. Protein-coding genes for photolyases and cryptochromes found in three genomes of UV-B-resistant bacteria isolated from HAAL.
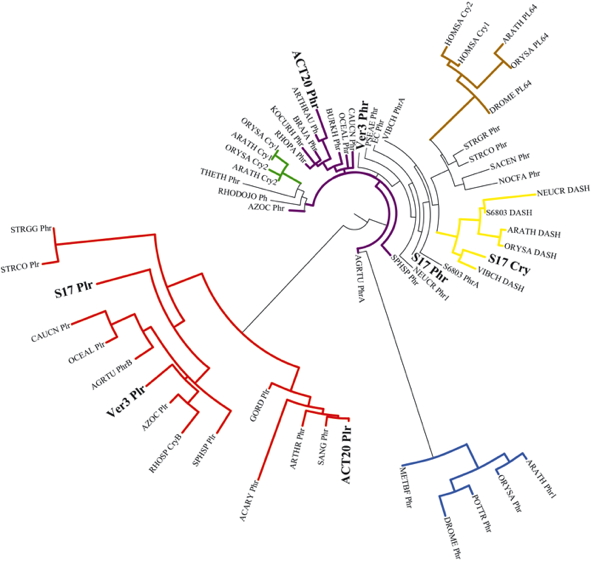
Figure 4. Phylogenetic tree of the cryptochrome/photolyase family. All sequences of photolyase-related proteins were retrieved from public databases via the National Center for Biotechnology Information web site. Multiple sequence alignments and the phylogenetic tree were carried out using the online available program phylogeny.fr (Dereeper et al., 2008). The species names are abbreviated as follows: Acaryochloris sp. CCMEE 5410 (Acary), Acinetobacter sp. Ver3 (Ver3), Agrobacterium tumefaciens (Agrtu), Arabidopsis thaliana (Arath), Arthrobacter aurescens (Arthrau) Arthrobacter chlorophenolicus (Arthr), Azorhizobium caulinodans ORS571 (Azoc), Bradyrhizobium japonicum USDA 110 (Braja), Burkholderia glumae (Burkh), Caulobacter crescentus (Caucn), Drosophila melanogaster (Drome), Escherichia coli (EC), Exiguobacterium sp. S17 (S17), Gordonia terrae (Gord), Homo sapiens (Homsa), Kocuria rhizophila (Kocurh), Methanosarcina barkeri (Metbf), Nesterenkonia sp. Act20 (Act20), Neurospora crassa (Neucr), Nocardia farcinica (Nocfa), Oceanocaulix alexandrii (Oceal), Oryza sativa (Orysa), Potorous tridactylus (Pottr), Pseudomonas aeruginosa (Pseae), Rhodobacter sphaeroides 2.4.1 (Rhosp), Rhodococcus jostii (Rhodojo), Rhodopseudomonas palustris (Rhopa), Saccharopolyspora erythraea (Sacen), Sanguibacter keddieii (Sang), Sphingomonas sp. SKA58 (Sphsp), Streptomyces coelicolor (Strco), Streptomyces griseus (Strgr), Streptomyces griseus subsp. griseus (Strgg), Synechocystis sp. PCC 6803 (S6803), Thermus thermophilus HB8 (Theth), and Vibrio cholerae (Vibch).
Groups within the CPF proteins are indicated in different colors: In green, Plant-Cry; in red, Cry-Pro and related iron-sulfur cluster containing bacterial cryptochromes and photolyases; in light brown, (6-4) photolyase and animal Cry; in yellow, DASH; in black, Class I CPD photolyases; in blue, Class II CPD photolyases; in purple, Class III CPD photolyases; Cry: cryptochrome, DASH: DASH cryptochrome, PL64: (6-4)photolyase, Phr: photolyase, Plr: photolyase-related protein, CPD: cyclobutane pyrimidine dimer, CryPro: proteobacteria and cyanobacteria cryptochromes.
In the genome of the planktonic strain Ver3, ORF sequences (PL1 and PL2) were found with identity to sequences within the CPF. On the protein sequence level, PL1 displayed a 64% identity or less with previous described CPD-Class I photolyases, while PL2 displayed 71% identity or less with different types of PLs called photolyase-related proteins (PRPs), now grouped according to Geisselbrecht et al. (2012) in the new clade Cry-Pro (Table 2). This affiliation was confirmed when a phylogenetic tree was constructed using HAAL Cry-PLs sequences, and the closest reference sequences in the database (Figure 4).
Inspection of the genome of the eu-endolithic strain S17 isolated from the stromatolites in Laguna Socompa (Belfiore et al., 2013) revealed three proteins belonging to the CPF family: S17-PL1 with a maximum of 74% identity with other Exiguobacterium photolyases, clustered also within the Class I CPD-photolyases and S17-PL2 with maximum of 82% identity with other Exiguobacterium PRP clustered near the group of PL-Cry, although in a different branch from Ver3. Another Cry was annotated in the genome of S17, and the comparison with the database revealed a 72% identity with the cry-DASH of other Exiguobacterium strains. Conversely, this protein gathered in the phylogenetic tree in the same cluster of cry-DASH proteins (Figure 4). The soil bacteria, Act20, also revealed two coding sequences with an identity with PLs: Act20-PL1 with a 55% or lower identity with CPD photolyases from actinobacterial strains, and Act20-PL2 with an identity with PLR proteins of actinobacterial strains of 67% or below. Interestingly, Act20-PL1 does not cluster within the group of the so-called actinomycetes CPD-photolyases, but is found in a sister branch of the plant cryptochromes and Class III CPD-photolyases.
The currently the best-characterized photolyase from HAAL is PL1 from Acinetobacter sp. Ver3 (Figure 4). Based on sequence alignments and secondary structure predictions for this novel photolyase and entries in the Protein Data Bank (PDB), we found the highest three-dimensional similarity to the photolyase from E. coli (PDB 1DNPA) (Albarracín et al., 2012). This photolyase belongs to the Class I CPD-photolyases, in which the well-known photolyase from E. coli is encountered (Park et al., 1995). Even as the sequence similarity of Ver3 to that of E. coli is low (41%), we found a high kinship between the 3D structures, as could be demonstrated from a model-building approach that indicated a nearly congruent protein folding (Figure 5).
The structure revealed a proximal alpha-beta domain, and a distal helical domain that binds to FAD, in full accordance to the structure of the E. coli photolyase. The N-terminal anti-parallel bundle of beta sheets enclosed by alpha helices is a typical folding motif of photolyases. The residues Asp105, Glu106, Lys298, Glu368, and Leu380 that interact with the antenna chromophore 5,10-methenyltetrahydrofolylpolyglutamate (MTHF) are fully conserved between the E. coli enzyme and the Ver3 photolyase, while in place of cys292 (E. coli protein) there is a serine residue (Ser297) in Ver3 photolyase. The residues Tyr225, Thr237, Ser238, Leu240, Ser241, Trp276, Arg283, Trp343, Asn346, Asp377, Asp379, and Ala382 that interact with FAD are all conserved in Ver3, except for Gln239, which replaces an arginine residue (Arg236) found in the E. coli photolyase (Albarracin et al., 2012). The chain of three tryptophan residues (Trp311, Trp364, and Trp387) instrumental for the electron transfer reaction (Brettel and Byrdin, 2010) is also conserved in Ver3 photolyase. The structural arrangement of this Trp-triade demonstrates the close proximity of these three residues to the isoalloxazine ring of FAD. This arrangement supports the putative CPD-photolyase property of this protein, which agrees with the efficient ability of Ver3 for repairing CPDs lesions.
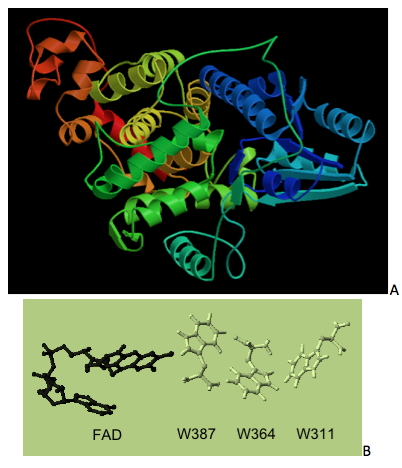
Figure 5. (A) Three-dimensional modelling of PL1, the CPD-Class I photolyase from Acinetobacter sp. Ver3, based on the three-dimensional structure of its E. coli ortholog (PDB 1DNPA). (B) Structural arrangement of FAD and three tryptophanresidues (that take part in electron transfer) in the 3D model structure from Ver3 photolyase (Albarracín et al., 2012).
Despite a high level of sequence diversity, the topological similarity between all photolyase structures known today is one of the most surprising discoveries. In Müller and Carell (2009), there is a comparison of the structures of 9 known cryptochromes and photolyases. Taking into account these structures, they calculated the total root mean square deviation for the amino acid residues that are in the proteins. This method compares the coordinates from the 3D structure for the backbone carbons (although the side chains adopt different orientations). The authors found that even when the sequences were different, the topology was not variable, resulting in deviations of only 1.914 Å over 369 residues, as calculated with Secondary-Structure Matching. It seems that all these proteins utilize similar energy and electron transfer steps, which, owing to the sensitivity of these processes, require a precise setting of distance and orientation. This apparently forces the proteins to maintain a common fold in order to ensure an optimal arrangement of the essential cofactors.
Concluding Remarks
The HAAL are natural laboratories for exploring and monitoring in situ interactions between the geophysical environment and the dynamics of biodiversity. Solar irradiation (including high UV-B doses) is without doubt the factor that puts the greatest pressure on the ecology of the microbial communities thriving in these shallow lakes. Thus, it is not surprising that ~100 strains with intrinsic UV-B-resistance have been isolated from HAAL. These organisms have developed strategies to cope with strong UV-B irradiation to avoid severe UV-B-damage to proteins, lipids and DNA. Hence, microbiota at the HAAL may harbor special mechanisms to sense and respond to such a ubiquitous resource, i.e., light. In accordance with this, we have encountered a rich diversity of photoreceptors within the cryptochrome-photolyase family in the genomes of three UV-B-resistant extremophiles. Of particular importance among these photoreceptors was the finding of a photolyase-coding sequence in the genome of Acinetobacter sp. Ver3 strain that displayed efficient photoreactivation ability after strong UV-B-induced DNA damage.
Based on sequence alignments and secondary structure predictions for this novel photolyase, and entries in the PDB, we found the highest three-dimensional similarity to the photolyase from E. coli (PDB 1DNPA). The structure revealed a proximal alpha-beta domain, and a distal helical domain that binds to FAD in full accordance to the structure of the E. coli photolyase. The N-terminal anti-parallel bundle of beta sheets enclosed by alpha helices is a typical folding motif of photolyases. The chain of three-tryptophan residues instrumental for electron transfer reaction is also conserved in Ver3 photolyase, and in close proximity to the isoalloxazine ring of FAD. All the former findings support the putative CPD-photolyase property of this protein, and agree with the efficient ability of Ver3 for repairing CPD lesions. Nevertheless, a more detailed functional characterization of this "extremoenzyme" is being conducted at the moment to clarify its repair function.
The scenario pictured herein makes the HAAL microorganisms excellent test cases for exploring novel enzymatic functions driven by light, and for the bioprospection of novel molecules with potential biotechnological applications on energy conversion, biomedicine or industry. This brief overview is intended to "shine" scientific light on a high window from a quite unexplored, exotic environment, which otherwise constitutes an exceptional outdoor photobiology lab.
References
Abdel-El-Haleem D (2003) Acinetobacter: Environmental and biotechnological applications. Afr J Biotechnol 2: 71-74.
Agogue H, Joux F, Obernosterer I, Lebaron P (2005) Resistance of marine bacterioneuston to solar radiation. Appl Environ Microbiol 71: 5282-5289.
Albarracín, V.H., Pathak, G., Douki, T., Cadet, J., Borsarelli, C., Gärtner, W., Farias, M.E. (2012) Extremophilic Acinetobacter strains from High-Altitude Lakes in Argentinean Puna: Remarkable UV-B Resistance and Efficient DNA Damage Repair. Origin of Life and Evolution of Biospheres. 42: 201-221.
Albarracín, V.H., Farias, M.E. (2012) Biotecnología Turquesa. Revista Hipótesis, Apuntes Científicos Uniandinos, Universidad de Los Andes, 13: 32-39.
Banaszak AT (2003) Photoprotective physiological and biochemical response of aquatic organisms to UVR. In UV effects in aquatic organisms and ecosystems. Helbling EW, Zagarese H (eds). Cambridge: The Royal Society of Chemistry, pp. 329-356.
Battista, J. R., A. M. Earl, and M. J. Park. (1999) Why is Deinococcus radiodurans so resistant to ionizing radiation? Trends Microbiol. 7:362–365.
Belfiore, Carolina; Ordoñez Omar, Farías María Eugenia (2013) Proteomic Approach of Adaptive Response to Arsenic Stress in Exiguobacterium sp. S17. An Extremophile Strain Isolate from High Altitude Andean Lake Stromatolite. Extremophiles, in press.
Bequer Urbano, S., Albarracín, V.H., Ordoñez, O.F., Farias, M.E., Alvarez, H.A. (2013) Lipid storage in High-Altitude Andean Lakes extremophiles and its mobilization under stress conditions in Rhodococcus sp. A5, a UV-resistant actinobacterium. Extremophiles, DOI 10.1007/s00792-012-0508-2.
Brettel K, Byrdin M (2010) Reaction mechanisms of DNA photolyase. Curr Opin Struct Biol 20: 693-701.
Buma AG, Boelen P, Jeffrey WH, Helbling E, Zagarese H (2003) UVR-induced DNA damage in aquatic organisms. In UV effects in aquatic organisms and ecosystems. Helbling EW, Zagarese H (eds). Cambridge: Royal Society of Chemistry, pp. 291-327.
Cabrol NA, Grin EA, Kiss KT, Ács E, Grigorszky I, Szabò K, Tóth B, Fike DA, Hock AN, Demergasso C, Escudero L, Chong G, Galleguillos P, Grigsby BH, Zambrana Román J, McKay CP, Tambley C. (2007) Signatures of Habitats and Life in Earth's High-Altitude Lakes: Clues to Noachian Aqueous Environments on Mars. In The Geology of Mars. Chapman M. (ed.). Cambridge University Press, pp. 349-370.
Cashmore AR (2003) Cryptochromes: enabling plants and animals to determine circadian time. Cell 114: 537–543.
Daly, M. J., L. Ouyang, P. Fuchs, and K. W. Minton. (1994) In vivo damage and recA-dependent repair of plasmid and chromosomal DNA in the radiation-resistant bacterium Deinococcus radiodurans. J. Bacteriol. 176:3508–3517.
Daly, M. J., and K. W. Minton. (1996) An alternative pathway of recombination of chromosomal fragments precedes recA-dependent recombination in the radioresistant bacterium Deinococcus radiodurans. J. Bacteriol. 178:4461–4471.
Dereeper A., Guignon V., Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.-F., Guindon S., Lefort V., Lescot M., Claverie J.-M., Gascuel O. Phylogeny.fr: robust phylogenetic analysis for the non-specialist Nucleic Acids Research. (2008) Jul 1; 36 (Web Server Issue):W465-9. Epub 2008 Apr 19. (PubMed)
Dib J, Motok J, Zenoff VF, Ordonez O, Farias ME (2008) Occurrence of resistance to antibiotics, UV-B, and arsenic in bacteria isolated from extreme environments in high-altitude (above 4400 m) andean wetlands. Curr Microbiol 56: 510-517.
Di Capua C, Bortolotti A, Farias, ME, Cortez, N (2011) UV-resistant Acinetobacter sp. isolates from Andean wetlands display high catalase activity. FEMS Microbiol Lett. 317: 181-189.
Farias ME, Fernandez-Zenoff V, Flores R, Ordonez O, Estevez C (2009) Impact of solar radiation on bacterioplankton in Laguna Vilama, a hypersaline Andean lake (4650 m). J Geophys Res 114: G00D04. doi:10.1029/2008JG000784
Farias, M.E., Rascovan, N., Toneatti, D.M., Albarracín, V.H., Flores, M.R., Poiré, D., Collavino, M., Aguilar, M., Vazquez, M., Polerecky, L. (2013) The discovery of stromatolites developing in a high-altitude volcanic lake Socompa, Argentinean Andes. PLoS ONE 8(1): e53497. doi:10.1371/journal.pone.0053497.
Farias, M.E., Poire, D.G., Arroiu, J., Albarracin, V.H. (2011) Modern stromatolite ecosystems at alkaline and hipersalyne high-altitude lakes at the Argentinean Puna. En V. Tewari y J. Seckbach (Eds.) Stromatolites: interaction of microbes with sediments. Cellular Origin, Life in Extreme Habitats and Astrobiology Book Series, 18: 427-441, Springer. DOI: 10.1007/978-94-007-0397-1. ISBN: 978-94-007-0396-4.
Fernandez-Zenoff V, Sineriz F, Farias ME (2006) Diverse Responses to UV-B Radiation and Repair Mechanisms of Bacteria Isolated from High-Altitude Aquatic Environments. Appl Environ Microbiol 72: 7857-7863.
Flores MR, Ordoñez OF, Farías ME (2009) Isolation of UV-B resistant bacteria from two high altitude Andean lakes (4,400 m) with saline and non saline conditions. J Gen Appl Microbiol 55: 447-458.
Fujihashi M, Numoto N, Kobayashi Y, Mizushima A, Tsujimura M, Nakamura A et al. (2007) Crystal Structure of Archaeal Photolyase from Sulfolobus tokodaii with Two FAD Molecules: Implication of a Novel Light-harvesting Cofactor. J Mol Biol 365: 903-910.
Haddad, A.F., Camacho, P., Durand, Cary S.C. (1995) Phylogenetic characterization of the epibiotic bacteria associated with the hydrothermal vent polychaete Alvinella pompejana. Appl. Environ. Microbiol. 61: 1679–1687.
Häder, D.P., Kumar, H.D., Smith R.C., Worrest R.C. (2007) Effects of solar UV radiation on aquatic ecosystems and interactions with climate change. Photochem Photobiol Sci 6: 267–285.
Halac, S., Felip, M., Camarero, L., Sommaruga-Wögrath, S., Psenner, R., Catalan, J., Sommaruga R. (1997) An in situ enclosure experiment to test the solar UV-B impact on microplankton in a high altitude mountain lake: lack of effect on phytoplankton species composition and growth. J Plankton Res 11: 1671–1687.
Hernandez KL, Quinones RA, Daneri G, Farias ME, Helbling EW (2007) Solar UV radiation modulates daily production and DNA damage of marine bacterioplankton from a productive upwelling zone (36o S), Chile. J Exp Mar Biol Ecol 343: 82-95.
Jiang, H., Dong, H., Yu, B., Liu, X., Li, Y., Ji, S., Zhang, C.L. (2007) Microbial response to salinity change in Lake Chaka, a hypersaline lake on Tibetan plateau. Environ. Microbiol. 10:2603–2621.
Joux F, Jeffrey WH, Lebaron P, Mitchell DL (1999) Marine bacterial isolates display diverse responses to UV-B radiation. Appl Environ Microbiol 65: 3820-3827.
Klar T, Kaiser G, Hennecke U, Carell T, Batschauer A, Essen LO (2006) Natural and non-natural antenna chromophores in the DNA photolyase from Thermus thermophilus. Chembiochem 7: 1798-1806.
Lange, C. C., L. P. Wackett, K. W. Minton, and M. J. Daly. (1998) Engineering a recombinant Deinococcus radiodurans for organopollutant degradation in radioactive mixed waste environments. Nat. Biotechnol. 16:929–933.
Liu, Y., Ya, T., Jiao, N., Kang, S., Zeng, Y., Huang, S. (2006) Microbial community structure in moraine lakes and glacial meltwaters, Mount Everest. FEMS Microbiol Lett 265(1):98–105.
Makarova KS, Aravind L, Wolf YI, Tatusov RL, Minton KW, Koonin EV, Daly MJ. (2001) Genome of the extremely radiation-resistant bacterium Deinococcus radiodurans viewed from the perspective of comparative genomics. Microbiol Mol Biol Rev. 65:44-79.
Mattimore, V., and J. R. Battista. (1996) Radioresistance of Deinococcus radiodurans: functions necessary to survive ionizing radiation are also necessary to survive prolonged desiccation. J. Bacteriol. 178:633–637.
Minton, K. W. (1994) DNA repair in the extremely radioresistant bacterium Deinococcus radiodurans. Mol. Microbiol. 13:9–15.
Müller M, Carell T (2009) Structural biology of DNA photolyases and cryptochromes. Curr Opin Struct Biol 19: 277-285.
Oberpichler I, Pierik AJ, Wesslowski J, Pokorny R, Rosen R, Vugman M, Zhang F, Neubauer O, Ron EZ, Batschauer A, Lamparter T. (2011) A photolyase-like protein from Agrobacterium tumefaciens with an iron-sulfur cluster. PLoS One. 6(10):e26775.
Ordonez OF, Flores MR, Dib JR, Paz A, Farias ME (2009) Extremophile Culture Collection from Andean Lakes: Extreme Pristine Environments that Host a Wide Diversity of Microorganisms with Tolerance to UV Radiation. Microb Ecol 58: 461-473.
Park HW, Kim ST, Sancar A, Deisenhofer J (1995) Crystal-Structure of DNA Photolyase from Escherichia coli. Science 268: 1866-1872.
Ponder, M..A, Gilmour, S.J., Bergholz, P.W., Mindock, C.A., Hollingsworth, R., Thomashow, M.F., Tiedje, J.M. (2005) Characterization of potential stress responses in ancient Siberian permafrost psychroactive bacteria. FEMS Microbiol Ecol 53:103–115.
Sancar A (2003) Structure and function of DNA photolyase and cryptochrome blue-light photoreceptors. Chem Rev 103: 2203-2237.
Sancar GB (2000) Enzymatic photoreactivation: 50 years and counting. Mutat Res, Fundam Mol Mech Mutagen 451: 25-37.
Sanchez LA, Gomez FF, Delgado OD (2009) Cold-adapted microorganisms as a source of new antimicrobials. Extremophiles 13: 111-120.
Seufferheld MJ, Alvarez HM, Farias ME (2008) Role of Polyphosphates in Microbial Adaptation to Extreme Environments. Appl Environ Microbiol 74: 5867-5874.
Sinha RP, Häder D-P (2002) UV-induced DNA damage and repair: a review. Photochem Photobiol Sci 1: 225-236.
Sweet, D. M., and B. E. Moseley. (1974) Accurate repair of ultraviolet induced damage in Micrococcus radiodurans. Mutat. Res. 23:311–318.
Sweet, D. M., and B. E. Moseley. (1976) The resistance of Micrococcus radiodurans to killing and mutation by agents which damage DNA. Mutat.Res. 34:175–186.
Tamada T, Kitadokoro K, Higuchi Y, Inaka K, Yasui A, deRuiter PE et al. (1997) Crystal structure of DNA photolyase from Anacystis nidulans. Nat Struct Biol 4: 887-891.
Towner KJ (2009) Acinetobacter: an old friend, but a new enemy. J Hosp Infect 73: 355-363.
Weber S (2005) Light-driven enzymatic catalysis of DNA repair: a review of recent biophysical studies on photolyase. Biochim Biophys Acta 1707: 1-23.
Williamson, Craig, E., Role, W. (1995) Does UV-B radiation play in freshwater ecosystems? Limnol Oceanogr 40: 386–392.
Winter C, Moeseneder MM, Herndl GJ (2001) Impact of UV radiation on bacterioplankton community composition. Appl Environ Microbiol 67: 665-672.
Zenoff V, Heredia J, Ferrero M, Sineriz F, Farias ME (2006) Diverse UV-B resistance of culturable bacterial community from high-altitude wetland water. Curr Microbiol 52: 359-362.
05/04/13